General Concepts
Structure and Function
Viruses are small obligate intracellular parasites, which by definition contain either a RNA or DNA genome surrounded by a protective, virus-coded protein coat. Viruses may be viewed as mobile genetic elements, most probably of cellular origin and characterized by a long co-evolution of virus and host. For propagation viruses depend on specialized host cells supplying the complex metabolic and biosynthetic machinery of eukaryotic or prokaryotic cells.
- A complete virus particle is called a virion.
- The main function of the virion is to deliver its DNA or RNA genome into the host cell so that the genome can be expressed (transcribed and translated) by the host cell.
- The viral genome, often with associated basic proteins, is packaged inside a symmetric protein capsid.
- The nucleic acid-associated protein, called nucleoprotein, together with the genome, forms the nucleocapsid. In enveloped viruses, the nucleocapsid is surrounded by a lipid bilayer derived from the modified host cell membrane and studded with an outer layer of virus envelope glycoproteins.
Classification of Viruses
Morphology: Viruses are grouped on the basis of
- size and shape,
- chemical composition and structure of the genome,
- and mode of replication.
Helical morphology is seen in nucleocapsids of many filamentous and pleomorphic viruses. Helical nucleocapsids consist of a helical array of capsid proteins (protomers) wrapped around a helical filament of nucleic acid. Icosahedral morphology is characteristic of the nucleocapsids of many "spherical" viruses. The number and arrangement of the capsomeres (morphologic subunits of the icosahedron) are useful in identification and classification. Many viruses also have an outer envelope.
Chemical Composition and Mode of Replication: The genome of a virus may consist of DNA or RNA, which may be
- single stranded (ss) or
- double stranded (ds),
- linear or circular.
- The entire genome may occupy either one nucleic acid molecule (monopartite genome) or several nucleic acid segments (multipartite genome). The different types of genome necessitate different replication strategies.
Nomenclature
Aside from physical data, genome structure and mode of replication are criteria applied in the classification and nomenclature of viruses, including the chemical composition and configuration of the nucleic acid, whether the genome is monopartite or multipartite. The genomic RNA strand of single-stranded RNA viruses is called sense (positive sense, plus sense) in orientation if it can serve as mRNA, and antisense (negative sense, minus sense) if a complementary strand synthesized by a viral RNA transcriptase serves as mRNA. Also considered in viral classification is the site of capsid assembly and, in enveloped viruses, the site of envelopment.
STRUCTURE AND FUNCTION
Viruses are inert outside the host cell. Small viruses, e.g., polio and tobacco mosaic virus, can even be crystallized. Viruses are unable to generate energy. As obligate intracellular parasites, during replication, they fully depend on the complicated biochemical machinery of eukaryotic or prokaryotic cells. The main purpose of a virus is to deliver its genome into the host cell to allow its expression (transcription and translation) by the host cell.
A fully assembled infectious virus is called a virion. The simplest virions consist of two basic components:
· nucleic acid (single- or double-stranded RNA or DNA)
· and a protein coat, the capsid, which functions as a shell to protect the viral genome from nucleases and which during infection attaches the virion to specific receptors exposed on the prospective host cell. Capsid proteins are coded for by the virus genome. Because of its limited size (Table 41-1) the genome codes for only a few structural proteins (besides non-structural regulatory proteins involved in virus replication). Capsids are formed as single or double protein shells and consist of only one or a few structural protein species. Therefore, multiple protein copies must self assemble to form the continuous three-dimensional capsid structure. Self assembly of virus capsids follows two basic patterns:
- helical symmetry, in which the protein subunits and the nucleic acid are arranged in a helix, and
- icosahedral symmetry, in which the protein subunits assemble into a symmetric shell that covers the nucleic acid-containing core.
Some virus families have an additional covering, called the envelope, which is usually derived in part from modified host cell membranes.
Viral envelopes consist of a lipid bilayer that closely surrounds a shell of virus-encoded membrane-associated proteins. The exterior of the bilayer is studded with virus-coded, glycosylated (trans-) membrane proteins. Therefore, enveloped viruses often exhibit a fringe of glycoprotein spikes or knobs, also called peplomers. In viruses that acquire their envelope by budding through the plasma or another intracellular cell membrane, the lipid composition of the viral envelope closely reflects that of the particular host membrane.
The outer capsid and the envelope proteins of viruses are glycosylated and important in determining the host range and antigenic composition of the virion. In addition to virus-specified envelope proteins, budding viruses carry also certain host cell proteins as integral constituents of the viral envelope. Virus envelopes can be considered an additional protective coat. Larger viruses often have a complex architecture consisting of both helical and isometric symmetries confined to different structural components. Small viruses, e.g., hepatitis B virus or the members of the picornavirus or parvovirus family, are orders of magnitude more resistant than are the larger complex viruses, e.g. members of the herpes or retrovirus families.
Classification of Viruses
Viruses are classified on the basis of morphology, chemical composition, and mode of replication. The viruses that infect humans are currently grouped into 21 families, reflecting only a small part of the spectrum of the multitude of different viruses whose host ranges extend from vertebrates to protozoa and from plants and fungi to bacteria.
Morphology
Helical Symmetry
In the replication of viruses with helical symmetry, identical protein subunits (protomers) self-assemble into a helical array surrounding the nucleic acid, which follows a similar spiral path. Such nucleocapsids form
- rigid,
- highly elongated rods
- or flexible filaments; in either case, details of the capsid structure are often discernible by electron microscopy. In addition to classification as flexible or rigid and as naked or enveloped,
- helical nucleocapsids are characterized by length, width, pitch of the helix, and number of protomers per helical turn. The most extensively studied helical virus is tobacco mosaic virus (Fig. 41-1). Many important structural features of this plant virus have been detected by x-ray diffraction studies. Figure 41-2 shows Sendai virus, an enveloped virus with helical nucleocapsid symmetry, a member of the paramyxovirus family (see Ch. 30).

FIGURE 41-2 Fragments of flexible helical nucleocapsids (NC) of Sendai virus, a paramyxovirus, are seen either within the protective envelope (E) or free, after rupture of the envelope. The intact nucleocapsid is about 1,000 nm long and 17 nm in diameter; its pitch (helical period) is about 5 nm. (x200,000) (courtesy of A. Kalica, National Institutes of Health.)
Icosahedral Symmetry
An icosahedron is a polyhedron having 20 equilateral triangular faces and 12 vertices (Fig. 41-3). Lines through opposite vertices define axes of fivefold rotational symmetry: all structural features of the polyhedron repeat five times within each 360° of rotation about any of the fivefold axes. Lines through the centers of opposite triangular faces form axes of threefold rotational symmetry; twofold rotational symmetry axes are formed by lines through midpoints of opposite edges. An icosaheron (polyhedral or spherical) with fivefold, threefold, and twofold axes of rotational symmetry (Fig. 41-3) is defined as having 532 symmetry (read as 5,3,2).
FIGURE 41-3 Icosahedral models seen, left to right, on fivefold, threefold, and twofold axes of rotational symmetry. These axes are perpendicular to the plane of the page and pass through the centers of each figure. Both polyhedral (upper) and spherical (lower) forms are represented by different virus families.
Viruses were first found to have 532 symmetry by x-ray diffraction studies and subsequently by electron microscopy with negative-staining techniques. In most icosahedral viruses, the protomers, i.e. the structural polypeptide chains, are arranged in oligomeric clusters called capsomeres, which are readily delineated by negative staining electron microscopy and form the closed capsid shell (Fig. 41-4 a/b). The arrangement of capsomeres into an icosahedral shell (compare Fig. 41-4 with the upper right model in Fig. 41-3) permits the classification of such viruses by capsomere number and pattern. This requires the identification of the nearest pair of vertex capsomeres (called penton: those through which the fivefold symmetry axes pass) and the distribution of capsomeres between them.
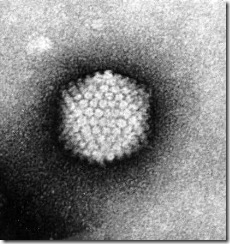
FIGURE 41-4a Adenovirus after negative stain electron microscopy. The capsid reveals the typical isometric shell made up from 20 equilateral triangular faces. The 252 capsomeres, 12 pentons and the 240 hollow hexon capsomeres are arranged in a T = 25 symmetry pattern vetite (x 400,000).
FIGURE 41-4b Adenovirus model. Capsomeres are depicted as circles surrounded by an electron dense stain. The inclined axes, h and k, are indicated. The second vertex has indices h = 5, k = 0. The total number of capsomeres C = 10(h2 + hk + k2) + 2 = 252. Capsomere organization is also expressed by the triangulation number, T, the number of unit triangles on each of the 20 faces of the icosahedron. A unit triangle is formed by lines joining the centers of three adjacent capsomeres. T = (h2 + hk + k2) = 25 for adenoviruses, and C = 1OT + 2. The 12 vertex capsomeres are surrounded by 5 other capsomeres each, therefore called penton and show 5-fold rotational symmetry. The penton base consists of 5 identical 85 kD polypeptide chains and extrudes a long antenna-like fiber protein. The 240 hexon capsomeres are trimers of the 120 kD hexon protomere polypeptide (for details see Ch. 67).
In the adenovirus model in Figure 41-4, one of the penton capsomeres is arbitrarily assigned the indices h = 0, k = 0 (origin), where h and k are the indicated axes of the inclined (60°) net of capsomeres. The net axes are formed by lines of the closest-packed neighboring capsomeres. In adenoviruses, the h and k axes also coincide with the edges of the triangular faces. Any second neighboring vertex capsomere has indices h = 5, k = 0 (or h = 0, k = 5). The capsomere number (C) can be determined to be 252 from the h and k indices and the equation: C = 10(h2 +hk + k2) + 2. This symmetry and number of capsomeres is typical of all members of the adenovirus family.
Virus Core Structure
Except in helical nucleocapsids, little is known about the packaging or organization of the viral genome within the core. Small virions are simple nucleocapsids containing 1 to 2 protein species. The larger viruses contain in a core the nucleic acid genome complexed with basic protein(s) and protected by a single- or double layered capsid (consisting of more than one species of protein) or by an envelope (Fig. 41-5).
FIGURE 41-5 Two-dimensional diagram of HIV-1 correlating (immuno-) electron microscopic findings with the recent nomenclature for the structural components in a 2-letter code and with the molecular weights of the virus structural (glyco-) proteins. SU stands for outer surface glycoprotein, TM for transmembrane gp, MA for membrane associated or matrix protein, LI for core-envelope-link, CA for major capsid, NC for nucleocapsid protein, respectively. PR, RT and IN represent the virus-coded enzymes protease, reverse transcriptase and integrase that are functional during the life cycle of a retrovirus (from Gelderblom, HR, AIDS 5, 1991).
Chemical Composition and Mode of Replication
RNA Virus Genomes
RNA viruses, comprising 70% of all viruses, vary remarkably in genome structure (Fig. 41-6). Because of the error rate of the enzymes involved in RNA replication, these viruses usually show much higher mutation rates than do the DNA viruses. Mutation rates of 10-4 lead to the continuous generation of virus variants which show great adaptability to new hosts. The viral RNA may be single-stranded (ss) or double-stranded (ds), and the genome may occupy a single RNA segment or be distributed on two or more separate segments (segmented genomes). In addition, the RNA strand of a single-stranded genome may be either a sense strand (plus strand), which can function as messenger RNA (mRNA), or an antisense strand (minus strand), which is complementary to the sense strand and cannot function as mRNA protein translation (see Ch. 42). Sense viral RNA alone can replicate if injected into cells, since it can function as mRNA and initiate translation of virus-encoded proteins. Antisense RNA, on the other hand, has no translational function and cannot per se produce viral components.
DsRNA viruses, e.g., members of the reovirus family, contain 10, 11 or 12 separate genome segments coding for 3 enzymes involved in RNA replication, 3 major capsid proteins and a number of smaller structural proteins. Each segment consists of a complementary sense and antisense strand that is hydrogen bonded into a linear ds molecule. The replication of these viruses is complex; only the sense RNA strands are released from the infecting virion to initiate replication.
The retrovirus genome comprises two identical, plus-sense ssRNA molecules, each monomer 7-11 kb in size, that are noncovalently linked over a short terminal region. Retroviruses contain 2 envelope proteins encoded by the env-gene, 4-6 nonglycosylated core proteins and 3 non-structural functional proteins (reverse transcriptase, integrase, protease: RT, IN, PR) specified by the gag-gene (Fig. 41-5).
- The RT transcribes the viral ssRNA into double-stranded, circular proviral DNA.
- This DNA, mediated by the viral integrase, becomes covalently bonded into the DNA of the host cell to make possible the subsequent transcription of the sense strands that eventually give rise to retrovirus progeny.
- After assembly and budding, retroviruses show structural and functional maturation. In immature virions the structural proteins of the core are present as a large precursor protein shell. After proteolytic processing by the viral protease the proteins of the mature virion are rearranged and form the dense isometric or cone-shaped core typical of the mature virion, and the particle becomes infectious.
DNA Virus Genomes
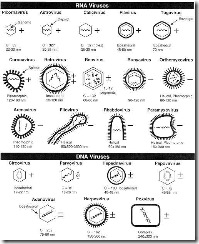
Most DNA viruses (Fig. 41-6) contain a single genome of linear dsDNA. The papovaviruses, comprising the polyoma- and papillomaviruses, however, have circular DNA genomes, about 5.1 and 7.8 kb pairs in size. DsDNA serves as a template both for mRNA and for self-transcription. Three or 2 structural proteins make up the papovavirus capsid: in addition, 5-6 nonstructural proteins are encoded that are functional in virus transcription, DNA replication and cell transformation.
FIGURE 41-6 Schemes of 21 virus families infecting humans showing a number of distinctive criteria: presence of an envelope or (double-) capsid and internal nucleic acid genome. +, Sense strand; -, antisense strand; ±, dsRNA or DNA; 0, circular DNA; C, number of capsomeres or holes, where known; nm, dimensions of capsid, or envelope when present; the hexagon designates the presence of an isometric or icosahedral outline.
Single-stranded linear DNA, 4-6 kb in size, is found with the members of the Parvovirus family that comprises the parvo-, the erythro- and the dependoviruses. The virion contains 2-4 structural protein species which are differently derived from the same gene product (see Ch. 64). The adeno-associated virus (AAV, a dependovirus) is incapable of producing progeny virions except in the presence of helper viruses (adenovirus or herpesvirus). It is therefore said to be replication defective.
Circular single-stranded DNA of only 1.7 to 2.3 kb is found in members of the Circovirus family which comprise the smallest autonomously propagated viruses. The isometric capsid measures 17 nm and is composed of 2 protein species only.
Virus Classification
On the basis of shared properties viruses are grouped at different hierarchical levels of order, family, subfamily, genus and species. More than 30,000 different virus isolates are known today and grouped in more than 3,600 species, in 164 genera and 71 families. Viral morphology provides the basis for grouping viruses into families. A virus family may consist of members that replicate only in vertebrates, only in invertebrates, only in plants, or only in bacteria. Certain families contain viruses that replicate in more than one of these hosts. This section concerns only the 21 families and genera of medical importance.
Besides physical properties, several factors pertaining to the mode of replication play a role in classification:
- the configuration of the nucleic acid (ss or ds, linear or circular),
- whether the genome consists of one molecule of nucleic acid or is segmented, and
- whether the strand of ss RNA is sense or antisense.
- Also considered in classification is the site of viral capsid assembly and,
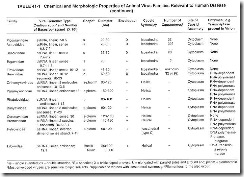
- in enveloped viruses, the site of nucleocapsid envelopment. Table 41-1 lists the major chemical and morphologic properties of the families of viruses that cause disease in humans.
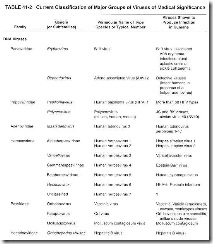
The use of Latinized names ending in -viridae for virus families and ending in -virus for viral genera has gained wide acceptance. The names of subfamilies end in -virinae. Vernacular names continue to be used to describe the viruses within a genus. In this text, Latinized endings for families and subfamilies usually are not used. Table 41-2 shows the current classification of medically significant viruses.
In the early days of virology, viruses were named according to common pathogenic properties, e.g. organ tropism and/or modes of transmission, and often also after their discoverers. From the early 1950s until the mid-1960s, when many new viruses were being discovered, it was popular to compose virus names by using sigla (abbreviations derived from a few or initial letters). Thus the name Picornaviridae is derived from pico (small) and RNA; the name Reoviridae is derived from respiratory, enteric, and orphan viruses because the agents were found in both respiratory and enteric specimens and were not related to other classified viruses; Papovaviridae is from papilloma, polyoma, and vacuolating agent (simian virus 40 [SV40]); retrovirus is from reverse transcriptase; Hepadnaviridae is from the replication of the virus in hepatocytes and their DNA genomes, as seen in hepatitis B virus. Hepatitis A virus is classified now in the family Picornaviridae, genus Hepatovirus. Although the current rules for nomenclature do not prohibit the introduction of new sigla, they require that the siglum be meaningful to workers in the field and be recognized by international study groups.
The names of the other families that contain viruses pathogenic for humans are derived as follows: Adenoviridae (adeno, "gland"; refers to the adenoid tissue from which the viruses were first isolated); Astroviridae (astron means star); Arenaviridae (arena "sand") describes the sandy appearance of the virion. Bunyaviridae (from Bunyamwera, the place in Africa where the type strain was isolated); Calicivirus (calix, "cup" or "goblet" from the cup-shaped depressions on the viral surfaces); Coronaviridae (corona, "crown") describes the appearance of the peplomers protruding from the viral surface; Filoviridae (from the Latin filum, "thread" or "filament") describes the morphology of these viruses. Herpesviridae (herpes, "creeping") describes the nature of the lesions; Orthomyxoviridae (ortho, "true," plus myxo "mucus," a substance for which the viruses have an affinity; Paramyxoviridae derived from para, "closely resembling" and myxo; Parvoviridae (parvus means, "small"); Poxviridae (pock means, "pustule"); Rhabdoviridae (rhabdo, "rod" describes the shape of the viruses and Togaviridae (toga, "cloak") refers to the tight viral envelope.
Several viruses of medical importance still remain unclassified. Some are difficult or impossible to propagate in standard laboratory host systems and thus cannot be obtained in sufficient quantity to permit more precise characterization. Hepatitis E virus, the Norwalk virus and similar agents (see Ch. 65) that cause nonbacterial gastroenteritis in humans are now assigned to the calicivirus family.
The fatal transmissible dementias in humans and other animals (scrapie in sheep and goat; bovine spongiform encephalopathy in cattle, transmissible mink encephalopathy; Kuru, Creutzfeldt-Jakob disease, and Gerstmann-Straussler-Scheinker syndrome in humans) (see Ch. 71 ) are caused by the accumulation of non-soluble amyloid fibrils in the central nervous systems. The agents causing transmissible subacute spongiform encephalopathies have been linked to viroids or virinos (i.e. plant pathogens consisting of naked, but very stable circular RNA molecules of about 3-400 bases in size, or infectious genomes enwrapped into a host cell coat) because of their resistance to chemical and physical agents. According to an alternative theory, the term "prion" has been coined to point to an essential nonviral infectious cause for these fatal encephalopathiesprion standing for self-replicating proteinaceous agent devoid of demonstrable nucleic acid. Some of the transmissible amyloidoses show a familial pattern and can be explained by defined mutations which render a primary soluble glycoprotein insoluble, which in turn leads to the pathognomonic accumulation of amyloid fibers and plaques. The pathogenesis of the sporadic amyloidoses, however, is still a matter of highly ambitious research.
REFERENCES
Caspar DLD: Design principles in virus particle construction. In Horsfall FL, Tamm I (eds): Viral and Rickettsial Infections in Man. 4th Ed. JB Lippincott, Philadelphia, 1975
Fields BN (ed): Virology. 3rd Ed. Lippincott-Raven Press, 1995
Gajdusek DC: Unconventional viruses and the origin and disappearance of kuru. Science 197:943, 1977
Gelderblom HR: Assembly and morphology of HIV: potential effect of structure on viral function. AIDS 5, 617-637,1991
Mattern CFT: Symmetry in virus architecture. In Nayak DP (ed): Molecular Biology of Animal Viruses. Marcel Dekker, New York, 1977
Morse SS (ed): The Evolutionary Biology of Viruses. Raven Press, New York, 1994
Murphy FA, Fauquet CM, Bishop DHL, et al. (eds): Virus Taxonomy: Sixth Report of the International Committee on Taxonomy of Viruses. Springer-Verlag, New York, 1995
Palmer EL, Martin ML: An Atlas of Mammalian Viruses. CRC Press, Boca Raton, 1988
Nermut MV, Stevens AC (eds): Animal Virus Structure. Elsevier, Amsterdam, 1989
Hans R. Gelderblom


 11.34
11.34
 Rafless bencoolen
Rafless bencoolen











0 komentar:
Posting Komentar